
Tamily Weissman-Unni
Associate professor of biology and department chair
Academic Credentials
PhD Columbia University 2004
BA Pomona College 1992
Teaching
Biology 202: Biology Core Concepts; Mechanisms
Biology/Psychology 252: Introduction to Neuroscience
Biology 422: Neurobiology with laboratory
Biology 490: Neuroanatomically Correct
Biology 490: Neural signaling & circuits
Research
Visit the lab website: weissmanlab.com
My lab is interested in the neural circuitry that underlies behavior. Specifically, how do connections develop in the young brain and how do mature connections generate behavior?
Brain function relies upon the precise organization of many neural circuits. Although we are beginning to understand important properties of simple circuits, we know little about how complex circuits form and how their mature properties underlie behavior. This is largely because the complexity of most brain circuits prevents their complete examination using traditional labeling or recording methods.
Due to the generation of a new powerful approach (“Brainbow”), we can now label populations of cells in many different colors, allowing us to visualize multiple individual components of a complex circuit in unprecedented detail. We have applied this approach to the translucent developing zebrafish nervous system, where individual cells can be visualized over time within the living animal. Using multicolor in vivo imaging approaches, we are testing how neural circuits form and function. The combination of two powerful approaches – Brainbow and zebrafish – allows us to ask these detailed questions about a complex circuit in the brain of a living, intact animal.
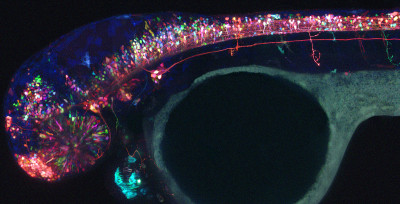
Our long-term goals are to understand how complex circuits develop and function in the brain. On a broader scale, we hope to shed light upon how the brain controls behavior and how this process can go awry in human disease. There are certain human disorders that arise when normal nervous system development goes wrong. A better understanding of normal and abnormal circuitry will help broaden our understanding of these diseases.
Publications (* indicates undergraduate author)
- Brockway NL, Cook ZC, *O’Gallagher MJ, Tobias ZJC, *Gedi M, *Carey KM, Unni VK, Pan YA, Metz MR & Weissman TA. Multicolor lineage tracing using in vivo time-lapse imaging reveals coordinated death of clonally related cells in the developing vertebrate brain. Developmental Biology, 2019 [in press]
- *Cook ZC, Brockway NL, Tobias ZJC, *Pajarla J, *Boardman IS, *Ippolito H, *Nkombo Nkoula S, & Weissman TA. Combining near-infrared fluorescence with Brainbow to visualize expression of specific genes within a multicolor context. Molecular Biology of the Cell, 2019 Feb 15;30(4):491-505. doi: 10.1091/mbc.E18-06-0340. Epub 2018 Dec 26. PMID: 30586321. Cover issue.
- Marra MH, *Tobias ZJC, *Cohen HR, Glover G, Weissman TA. In vivo time-lapse imaging in the zebrafish lateral line: A flexible, open-ended research project for an undergraduate neurobiology laboratory course. Journal for Undergraduate Neuroscience Education, 2015 Jul 7;13(3):A215-24. eCollection 2015 Summer.
- Weissman TA, Pan YA. Brainbow: New resources & emerging biological applications for multicolor genetic labeling & analysis. Genetics, 2015 Feb; 199(2):293-306. Cover issue.
- *Hamling KR, *Tobias ZJC, Weissman TA. Mapping the development of cerebellar Purkinje cells in zebrafish. Developmental Neurobiology, 2015 Feb 4 doi: 10.1002/dneu.22275
- Pan YA, Freundlich T, Weissman TA, Schoppik D, Wang CX, Ciruna B, Sanes JR, Lichtman JW, Schier AF. Zebrabow: multispectral cell labeling for cell tracing and lineage analysis in zebrafish. Development, 2013 July;140(13):2835-46.
- Weissman TA, Sanes JR, Lichtman JW, Livet J. Generating and Imaging Multicolor Brainbow Mice. Cold Spring Harbor Protocols, July 2011 (7:763-9).
- Livet J, Weissman TA, Kang H, Draft, RW, Lu, J, Bennis RA, Sanes JR, Lichtman JW. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature, Nov 1, 2007, 450 (7166):56-62.
- Unni VK, Weissman TA, Rockenstein E, Masliah E, McLean PJ, Hyman BT. In vivo imaging of a-synuclein in mouse cortex demonstrates stable expression and differential subcellular compartment mobility. PLoS ONE, May 2010, 5(5):e10589.
- Weissman T, Ivic L, Riquelme, PA, Flint AC, & Kriegstein AR. Calcium waves propagate through radial glial cells and modulate proliferation in developing cortex. Neuron, September 2, 2004, 43(5):647-661.
- Weissman T*, Noctor SC*, Clinton BK, Honig LS, & Kriegstein AR. Neurogenic radial glial cells in reptile, rodent, and human: from mitosis to migration. Cerebral Cortex, June 2003, 13(6):550-9.
- Labrakakis C, Tong C-K, Weissman T, Torsney C, and MacDermott AB. Localization and function of ATP and GABAA receptors expressed by nociceptors and other postnatal rat sensory neurons. Journal of Physiology, May 15, 2003, 549(Pt 1):131-42.
- Noctor SC, Flint AC, Weissman TA, Wong WS, Clinton BK, & Kriegstein AR. Dividing precursor cells of the embryonic cortical ventricular zone have morphological and molecular characteristics of radial glia. Journal of Neuroscience, April 15, 2002, 22(8):3161-3173.
- Noctor SC, Flint AC, Weissman TA, Dammerman RS, & Kriegstein AR. Neurons derived from radial glial cells establish radial units in neocortex. Nature, February 8, 2001, 409:714-720.
- Angelastro JM, Klimaschewski L, Tang, S, Vitolo OV, Weissman TA, Donlin LT, Shelanski ML, & Greene LA. Identification of diverse nerve growth factor-regulated genes by Serial Analysis of Gene profiling. PNAS, September 12, 2000, 97: 10424–10429.
- Naeser M, Baker E, Palumbo C, Nicholas M, Alexander M, Samaraweera R, M. Prete M, Hodge S, & Weissman T. Lesion site patterns in severe nonverbal aphasia to predict outcome with a computer- assisted treatment program. Archives of Neurology, November, 1998, 55(11):1438-48.
Location: Biology-Psychology Hall
Biology is located in room 210 of Biology-Psychology on the Undergraduate Campus.
MSC: 53
email biology@lclark.edu
voice 503-768-7511
Chair Tamily Weissman-Unni
Biology
Lewis & Clark
615 S. Palatine Hill Road MSC 53
Portland OR 97219